At present, monoclonal antibodies have developed rapidly as therapeutic antibodies, but their clinical application is limited by some problems, such as inducing human anti-mouse antibody response, low tumor penetration, and short half-life. Phage display technology can prepare small molecule single chain antibodies, which makes it possible to produce fully humanized antibodies, and has gradually become one of the main means to obtain humanized antibodies.
Phage display technology is an efficient gene expression screening technology, that fuses foreign proteins or peptides with specific phage capsid proteins, displays them on the phage surface, and maintains relatively independent spatial conformation and biological activity, which is conducive to the specific recognition and binding of target molecules, to realize the unity of genotype and expression type[1]. In 1985, Smith first reported the invention of phage display technology using genetic engineering technology. So far, this technology has developed quite maturely and has been widely used in scientific research fields such as antigen epitope analysis, monoclonal antibody production, antibody humanization, drug screening, vaccine development, immunological disease diagnosis, and treatment.
Principle of Phage Antibody Display Technology
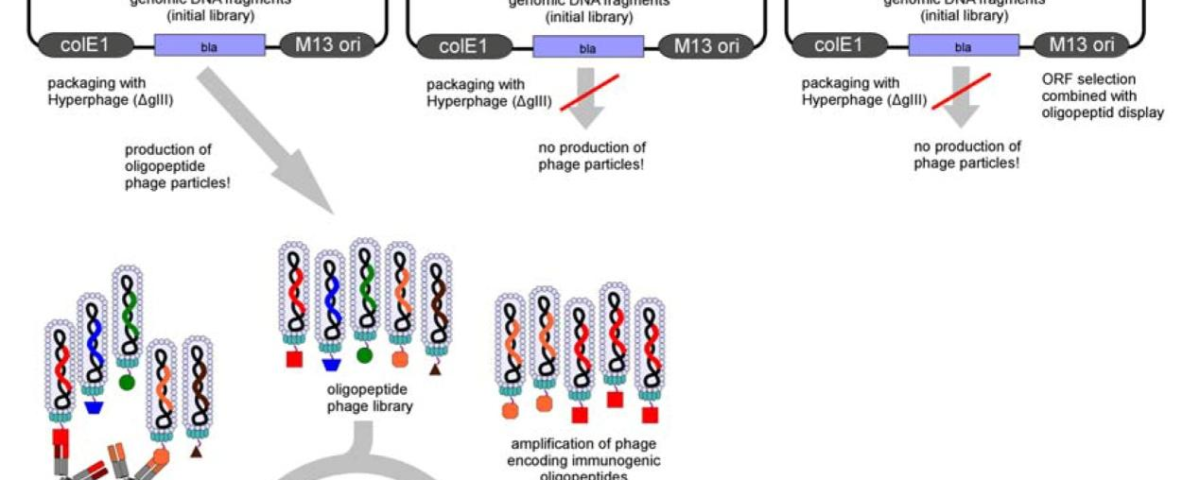
Phage display technology inserts the gene encoding exogenous peptides or proteins into the appropriate position of the structure gene of the phage coat protein through genetic engineering technology, so that the gene can be correctly expressed in the reading frame, so that the exogenous peptides or proteins can form fusion proteins on the capsid protein of the phage and be presented on the surface of the phage with the reassembly of the progeny phage. It can maintain relative spatial structure and biological activity. Then the target molecules were used to wash away the unspecifically bound bacteriophages by appropriate panning methods. Then the bound phage is eluted with acid-base or competing molecules, and the neutralized phage infects E. coli with amplification. After 3-5 rounds of enrichment, the proportion of phages that can specifically recognize target molecules is gradually increased, and finally, the polypeptide or protein that can recognize target molecules is obtained[2].
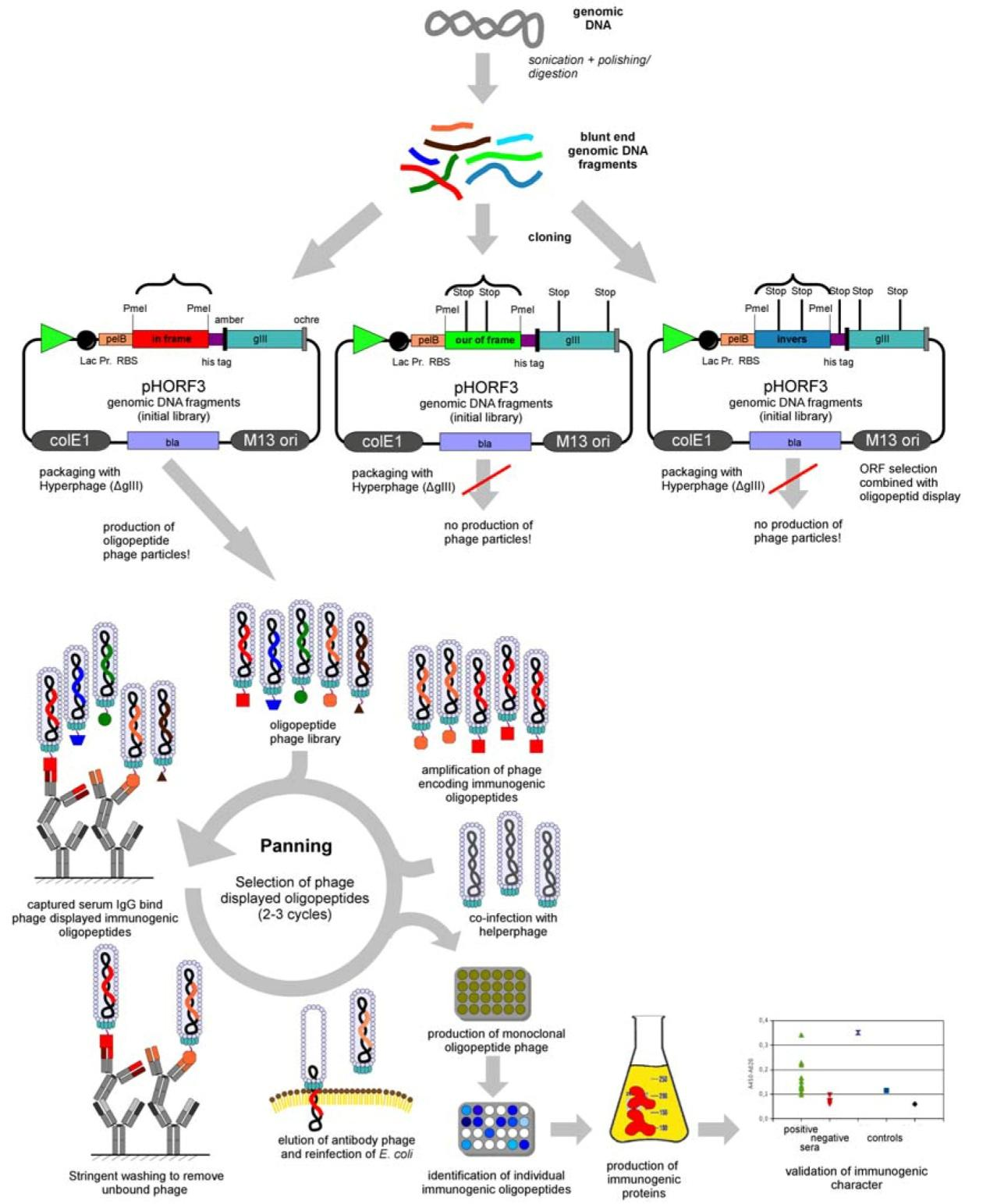
Figure 1:Flowchart of bacteriophage antibody library production(https://www.kmdbioscience.com/pages/phage-display-peptide-library-platform.html)
Classification of Phage Library
Phage library building refers to the H chain and L chain gene library of antibodies amplified from B lymphocytes, and the antibody fragment group is fused with phage capsid protein and displayed on the phage surface to form a phage display antibody library[1]. There are three main types: the scFv library (composed of VH and VL and joined by a small peptide (Gly4Ser) 3 into a single-stranded polypeptide), the Fab library (containing VH-CH, VL-Cl and joined together by disulfide bonds), and the VHH library (only heavy chain variable regions).
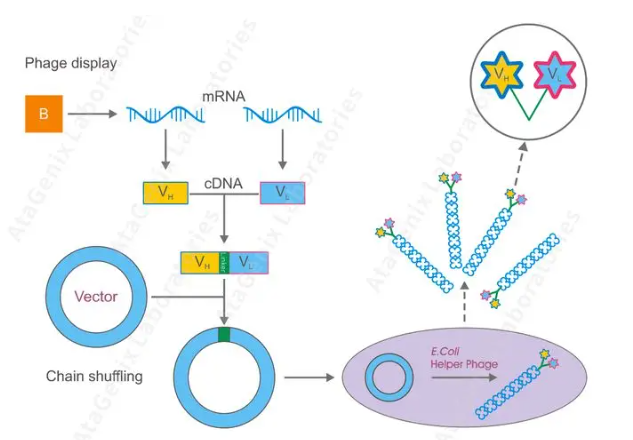
Figure 2: Phage Library Construction
Random Peptide Library
The random oligonucleotide sequence synthesized by chemical synthesis was fused with the phage surface protein gene, and the random sequence short peptides of various amino acid combinations were expressed on the phage surface. The phage display random peptide library can be used to screen the epitopes of various antigens in high throughput.
The phage display random peptide library can be divided into hexapeptide library, nine-peptide library, and twelve-peptide library according to different sizes. According to the structure, it can be divided into linear and nonlinear peptide libraries. According to the special purpose, it can be divided into a partial peptide library and a mutant peptide library.
Immune Bank
The immune bank is constructed from antibody genes of donor B lymphocytes after immunization (including vaccine injection, microbial infection, autoimmune disease, tumor, etc.). The efficiency of antibody panning for specific immune antigens is high, but it is generally only suitable for the selection of a specific antibody, and the storage capacity is not high, and the storage capacity of 106-108 can meet the needs. Compared with the natural library, the antibody with high affinity and high specificity can be screened from the immune library. Due to immune selection pressure, the abundance of specific antibodies against the corresponding immunogen in the library is much higher than that of other non-specific antibodies, some of which have undergone affinity maturation process of the immune system, and functional antibodies against the immunogen and their genes can be screened from libraries with small library capacity.
Natural Reservoir
Don’t immunize animals. The antibodies are all derived from humans, completely humanized antibodies, but the affinity is usually low. All antibody genes in and outside the bone marrow are included, and the antibody library constructed is suitable for the selection of antibodies corresponding to all antigens. The capacity of the library is relatively high, and the minimum need is 10^9 because the current estimated types of natural antigens are in this order of magnitude.
References
[1] Saw PE, Song EW. Phage display screening of therapeutic peptide for cancer targeting and therapy. Protein Cell. 2019 Nov;10(11):787-807. doi: 10.1007/s13238-019-0639-7. Epub 2019 May 28. PMID: 31140150; PMCID: PMC6834755.
[2] Juen L, Brachet-Botineau M, Parmenon C, Bourgeais J, Hérault O, Gouilleux F, Viaud-Massuard MC, Prié G. New Inhibitor Targeting Signal Transducer and Activator of Transcription 5 (STAT5) Signaling in Myeloid Leukemias. J Med Chem. 2017 Jul 27;60(14):6119-6136. doi: 10.1021/acs.jmedchem.7b00369. Epub 2017 Jul 12. PMID: 28654259.
[3] Potocnakova L, Comor L, Schreterova E,et al. B-cell mimotope mapping of anti-Borrelia bavariensis antibodies by a phage display peptide library[C]//ISMD2016 Eleventh International Symposium on Molecular Diagnostics.2016.