Immunoprecipitation (IP) is a widely employed technique in molecular biology and biochemistry, enabling the isolation and concentration of specific proteins from complex mixtures. By leveraging the specificity of antibodies to selectively bind proteins, IP facilitates the study of protein-protein interactions, protein modifications, and protein functions. This article delves into the myriad applications of immunoprecipitation and outlines some common protocols utilized in this technique.
The essence of immunoprecipitation lies in the interaction between the antigen (target protein) and the antibody. A suitable antibody is chosen to selectively bind the protein of interest, forming an antigen-antibody complex. This complex is then precipitated from the solution, typically using protein A/G agarose beads. The beads are collected, and washed to remove unbound materials, and the bound proteins are eluted for further analysis. Understanding the core principles and methodology of IP is pivotal for obtaining reliable and interpretable results.
Immunoprecipitation (IP) and other protein interaction applications such as yeast two-hybrid (Y2H), protein microarrays, and Förster Resonance Energy Transfer (FRET) serve the common goal of elucidating protein interactions, albeit through distinct mechanisms. IP leverages the specificity of antibodies to isolate and concentrate particular proteins from a mixture, providing a direct method of assessing protein interactions or modifications in a near-native state. On the other hand, Y2H employs a genetic system in yeast to detect interactions, offering a high-throughput albeit less direct method, which may miss interactions occurring outside the yeast’s nuclear environment. Protein microarrays provide a high-throughput platform to study protein-protein interactions on a large scale, yet may not always reflect the natural cellular context. FRET utilizes energy transfer between two light-sensitive molecules to gauge the interaction and distance between proteins, offering real-time interaction analysis within living cells. Each of these methods has its own set of advantages and limitations concerning sensitivity, throughput, and contextual relevance, choosing method contingent on the specific requirements of the research question at hand.
Immunoprecipitation finds extensive applications in research fields like oncology, immunology, and neurobiology. It is instrumental in studying protein-protein interactions, identifying post-translational modifications, and investigating cellular signaling pathways. Moreover, IP serves as a crucial step in various downstream applications such as Western blotting, mass spectrometry, and enzyme activity assays which provide insights into the functional and structural aspects of proteins.
Here’s a brief protocol of Immunoprecipitation:
Step 1 Sample Preparation
Preparing a clear, homogenous sample is the first step, which involves lysing cells or tissues to release proteins. Adding protease and phosphatase inhibitors to prevent protein degradation.
Step 2 Antibody Selection and Binding
Choosing a high-affinity, specific antibody is crucial. The antibody is incubated with the sample to allow binding to the target protein. Incubate at an appropriate temperature for a specified time (e.g., 1-2 hours) to allow antibody-protein binding.
Step 3: Immobilization to a Solid Support
The antibody-protein complexes are then immobilized on a solid support, usually beads coated with protein A or G which bind the Fc region of antibodies. During this incubation, the antibody binds to its target protein, and the protein-bead-antibody complex forms.
Step 4 Washing and Elution
Washing the beads several times with the wash buffer to remove unbound proteins and contaminants, and the target protein is eluted from the beads, often by altering the pH or using a competitive ligand.
Step 5 Analysis
The eluted protein can be analyzed using various techniques like Western blotting, mass spectrometry, or gel electrophoresis.
Step 6 Optimization
Fine-tuning parameters such as incubation times, temperatures, and buffer compositions is often necessary to achieve optimal results.
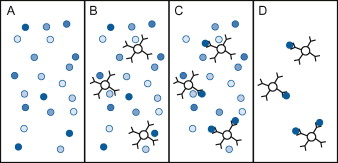
Figure 1. Immunoprecipitation. (A) Before an immunoprecipitation procedure begins, a protein of interest exists in a heterogenous sample with many other proteins. (B) A scientist adds antibodies that recognize the protein to small beads and then adds the beads to the sample. (C) The antibodies bind to the proteins, causing them to precipitate out of solution and adhere to the beads. (D) The beads are removed from the sample, purifying the protein of interest. The proteins can be removed from the beads by changing the salinity or pH of the solution.(Figure source: Matt Carter, Jennifer C. Shieh, 2010)
Immunoprecipitation remains a cornerstone technique, bridging the molecular interactions with the vast expanse of research insights. The meticulous adherence to protocols coupled with a profound understanding of its applications propels the scientific inquiry forward, opening avenues for novel discoveries and a deeper understanding of life’s molecular machinery.
KMD Bioscience has been dedicated to the study of protein-nucleic acid interactions for over 10 years. Eukaryotic genomic DNA exists in the form of chromatin, and the study of protein-DNA interactions in the chromatin environment is a fundamental way to elucidate the mechanisms of eukaryotic gene expression. Scientists in KMD Bioscience, which have accumulated a wealth of experience in immunoassay technology, are able to quickly customize protein-nucleic acid interaction study assays for different clients. With years of technical expertise, KMD Bioscience is not only able to complete the experimental content quickly, but also to ensure that each step of the experiment is carefully controlled to ensure that the test results are accurate, objective and credible. KMD Bioscience, which has complete protein detection equipment, has established mature and perfect antibody platforms, protein platforms and other technology platforms.
For more information, Visit us at https://www.kmdbioscience.com/ to have a detailed understanding.
