The diversity of antibodies in the body plays a highly effective role in defending against foreign pathogens. Although antibodies can be used as potential therapies, there are so many of them that only a fraction of them can currently be tested. In the past, flow sorting was usually used to identify B cells that produce specific antibodies to screen for immortal B cells. But these methods, which match functional screening to heavy – and light-chain gene sequences, are less efficient: typically, a B cell can be screened against only a few antigens simultaneously. With the development of second-generation sequencing (NGS) methods, high-throughput screening has improved the efficiency of studying antibody sequences. High-throughput screening systems automate the parallel analysis of thousands of reactions, increasing the experimental scale and efficiency of drug screening. Among them, cell-based high-throughput drug screening becomes the main screening mode because it can provide a microenvironment similar to the physiological conditions of the human body. However, matching antibody sequencing to its antigen requires the synthesis and identification of single-cell clonal antibodies. Recent research still combines antibody sequencing and function to solve the problem of high-throughput screening, although some techniques have facilitated the discovery of some neutralizing antibodies, but still face the problem of screening multiple antigens for a single B cell[1].
Dr. Ivelin S. Georgiev’s team from Vanderbilt University Medical Center in the United States, together with 17 other research institutions, Including a collaboration between Duke University School of Medicine and Harvard Medicine for a Cell paper on High-Throughput Mapping of B Cell ReceptorSequences to Antigen Specificity, they developed Libro-SEq linking B cell receptor to antigen specification through sequencing (linking B cell receptor to antigen specification through sequencing) method. HIV – and influenza-specific B cells were identified in two HIV-infected individuals, while the antigen response of thousands of individual B cells was predicted, and a new HIV-neutralizing antibody was discovered. This study provides a new way for antibody discovery[2].
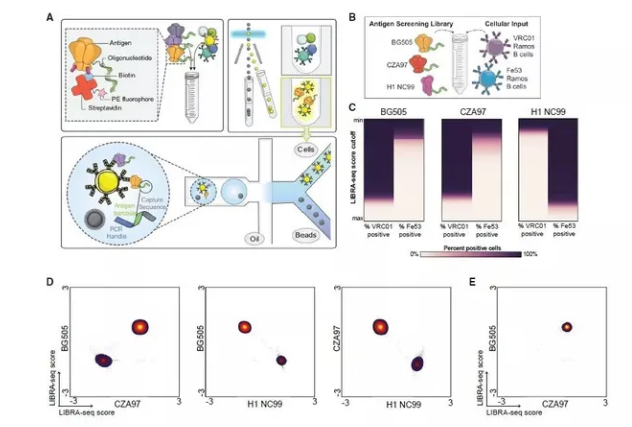
Figure 1: LIBRA-seq Assay Schematic and Validation[2]
Using Lip-SEQ as a second-generation sequence-based reading method applied to peripheral blood mononuclear cell samples from HIV-infected patients, the researchers successfully identified antibodies to HIV and influenza, including known and new neutralizing antibody lineages. After testing for accuracy with known B-cell receptors, they applied the technique to antibody discovery, confirming HIV-specific bNab in two known human samples and a new lineage of bNab. In addition, they found more HIV-specific antibodies and identified a class of them. Influenza-specific antibodies were isolated from HIV-1-infected patients by HA probe screening. These results indicate that the combination of the antibody sequence and its specificity can simultaneously screen out millions of B cells that can bind to specific antigens, avoiding the limitations of conventional fluorescence sorting. In addition to antibody discovery applications, Lilibra seq can also be applied to immune status monitoring, for example, they found that in B cells, B cells with a high score for several HIV-1 antigens LiLibra seq than B cells with a high score for only one HIV-1 antigen, their IGHV somatic mutations increased.
This study has developed and confirmed that LiP-SEQ is a high-throughput, measurable and promising technology. This method can profile single-cell clonal antibody sequences with a variety of different antigens, and can be applied to antibody discovery and immune status analysis, which is of great significance for the discovery of antibodies and vaccines. With the continuous accumulation of data, it will play a huge role in promoting the basic research and application of immunology.
References
[1] Busse, C.E., Czogiel, I., Braun, P., Arndt, P.F. & Wardemann, H. Single-cell based high-throughput sequencing of full-length immunoglobulin heavy and light chain genes. Eur J Immunol 44, 597-603 (2014).
[2] Setliff I, Shiakolas AR, Pilewski KA et al. High-ThroughputMapping of B Cell Receptor Sequences to Antigen Specificity Cell. 2019 Dec12;179(7):1636-1646.e15.

