Monoclonal antibodies are one of the most important types of biological drugs on the pharmaceutical market and can also be used for diagnostic purposes. The FDA has approved over 100 monoclonal antibodies for treating a wide range of diseases, including cancer, infectious diseases, autoimmune diseases, and neurological disorders. However, in the 10 years before 2020, the FDA has not approved any antibody drugs prepared by single B cell antibody production technology, and some shortcomings still need to be overcome and improved.
Monoclonal antibodies are often selected from antigen-specific single B cells derived from different hosts, which are notably short-lived in ex vivo culture conditions and hence, arduous to interrogate. The development of several new techniques and protocols has facilitated the isolation and retrieval of antibody-coding sequences of antigen-specific B cells by also leveraging miniaturization of reaction volumes. Alternatively, mAbs can be generated independently of antigen-specific B cells, comprising display technologies and, more recently, artificial intelligence-driven algorithms. Consequently, a considerable variety of techniques are used, raising the demand for better consolidation[1].
A rather novel methodology to identify antigen-specific B cells makes use of the recently developed single cell transcriptomic techniques, such as the 10x Chromium, or any system using beads carrying barcoded-oligos, such as Drop-Seq[2]. The concept is based on the fact that upon immunization or infection, among the most expanded clones (defined as a unique homologous combination of natively paired VH–VL), there is a higher prevalence of antigen-specific B cells[3]. Using conventional bulk Ig NGS techniques would be unable to directly deliver the genetic information of native antigen-specific monoclonal antibodies, because of the technical inability to retrieve natively paired individual VH–VL sequences. Despite this latter limitation, a first proof-of-principle study demonstrated the concept of antigen-specific BCR mining by using B cell sequencing data (not natively paired VH–VL sequences), analyzing properties such as clonal expansion, public clones, and mutational burden. The concept evolved further and a first panel of Ebola-virus-specific mAbs was identified based on the analysis of the most expanded clones upon Ebola VLP-based immunization of BALB/c mice and retrieval of natively paired VH–VL sequences using the picoliter well in a PDMS chip hosting individual B cells. The most expanded unique VH–VL sequences were chosen as candidates and then synthesized, cloned into suitable mammalian expression vectors, and expressed as recombinant mAbs to corroborate their antigen specificity (Figure 1).
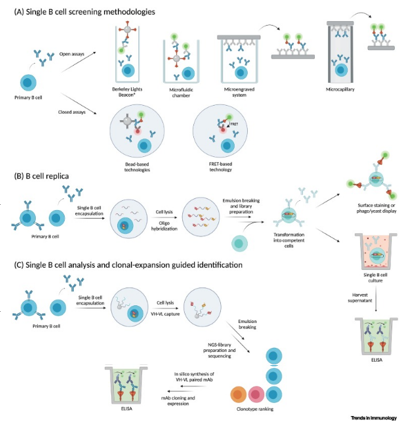
Figure 1. Single B cell screening technologies using miniaturized equipment[1].
Despite the advent of many in vitro display technologies, the development of mAbs also relies on ex vivo single B cell interrogation. Among the plethora of techniques enabling this process, the hybridoma technology is still used in new discovery pipelines because it is easy, cheap, and familiar to many scientists . The main problems lie on the traditionally difficult-to-overcome mindset of having to adapt to new techniques, and the technical difficulty and execution of the new protocols (if described at all), as well as the price tag of the new instruments, making these methodologies out of reach for many laboratories.
Despite their difficult implementation, major advances are provided by the emerging new technologies for the interrogation of single B cells using high-throughput screening approaches, especially for the identification of mAbs that are specific for ‘high-hanging fruit’ antigens such as GPCRs, ion channels, and intracellular targets. In addition to high-throughput systems, it will be crucial to implement efficient functional screening approaches, including for viruses and toxins, as well as for protein neutralization or receptor blockade (see Outstanding questions).
Furthermore, it is likely that when (and not if) artificial intelligence (AI)-based solutions are readily available, many of the antibody-discovery campaigns might be conducted almost completely in silico, with the important exception of the ensuing in vitro and in vivo validation. Especially in the last 2 years, major advances have been made in the field of AI-based in silico prediction of protein structures, which might allow to more precisely predict the structural basis of antibody–antigen interactions in the near future, without the need of wet-laboratory techniques a priori . However, we expect that further advancement of single B cell technologies will still remain highly significant in mAb discovery; indeed, research focused on the isolation of natively paired VH–VL mAbs or on target-agnostic screenings remain fundamental research areas in immunology .
References
[1] Pedrioli A , Oxenius A .Single B cell technologies for monoclonal antibody discovery[J].Trends in Immunology, 2021, 42(12):1143-1158.DOI:10.1016/j.it.2021.10.008.
[2] Macosko E Z , Basu A , Satija R ,et al.Highly Parallel Genome-wide Expression Profiling of Individual Cells Using Nanoliter Droplets[J].Cell, 2015, 161(5):1202-1214.DOI:10.1016/j.cell.2015.05.002.
[3] Truck J , Ramasamy M N , Galson J D ,et al.Identification of antigen-specific B cell receptor sequences using public repertoire analysis.[J].The Journal of Immunology, 2015, 194(1):252-261.DOI:10.4049/jimmunol.1401405.
